Integration Across This Project
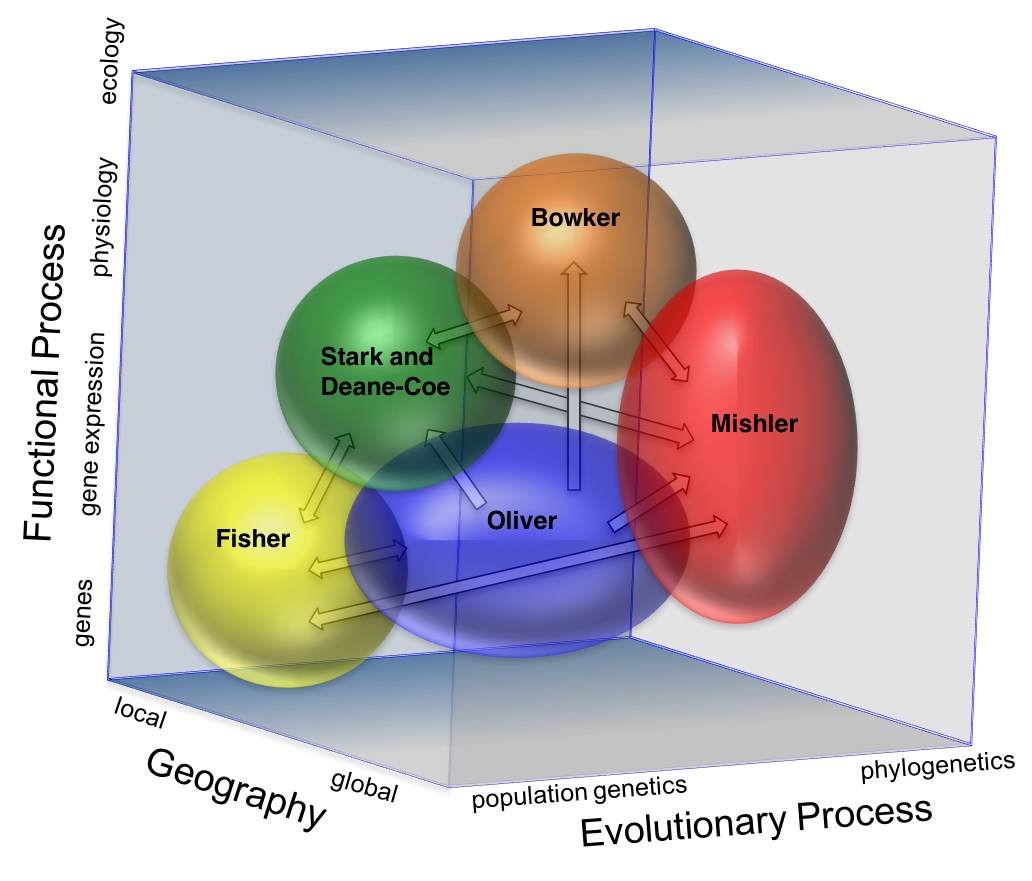
Our research program examines the past, present, and future drivers of biodiversity in this ecologically important dryland moss clade. Each lab has its own specific role to play, yet the various studies proposed above, in genetic, phylogenetic, and functional biodiversity, fit into an integrated conceptual and organizational model as visualized in Fig. 6. As shown in this 3-dimensional space, the different labs will concentrate their efforts at different scales of functional, geographical, and evolutionary processes, yet interact as described below to build up a synthetic picture of variation in desiccation tolerance and reproductive biology and the many factors that affect them. This will reveal powerful connections among different dimension of biodiversity, and the purported drivers of diversity in these mosses: desiccation tolerance and reproductive strategy.
Evolutionary process
At the broadest level, we will examine macroevolutionary patterns and processes that reveal phylogenetic relationships among species sampled across a large geographic range, create a robust phylogeny and taxonomic classification for Syntrichia, and use it to study trait evolution (Mishler lab). We will also examine the relationship between phylogenetic diversity and ecological function in Syntrichia in soil biocrust communities to understand the mechanistic drivers of phylogenetic distributions in a complex ecological landscape (Bowker lab). At a finer scale, we will focus on S. caninervis and S. ruralis, to examine the genomic/genetic (Oliver lab) and physiological (Stark and Deane-Coe labs) drivers of DT and variability in reproductive biology between species, and among populations within each species (Fisher lab).
Functional process
Patterns of biodiversity that manifest at the ecological level ultimately have a genetic basis, and our research program is designed to hierarchically address the functional determinants of traits that directly influence biodiversity. First, we will construct draft genomes for S. caninervis and S. ruralis, and then use transcriptomics to identify patterns of differential gene expression associated with environmental stress and developmental state in these focal species complexes (Oliver lab). Next, using the same populations, we will directly address physiological tradeoffs in DT and reproductive biology (sexual and asexual) among and within species by subjecting male and female shoots to desiccation stress treatments in the laboratory (Stark and Deane-Coe labs).
Moving to the ecological level, we will examine the relationship between genetic diversity and differential physiological stress in S. caninervis and S. ruralis, and biocrust community resilience to environmental change (Bowker lab). This integrated analysis of hierarchical functional relationships from genes to ecosystems allows us to directly infer past and current structural drivers of biodiversity as well as how patterns of diversity may change in a future climate. By carefully tracking genotypic variation and phenotypic plasticity at all levels of the study it is possible that we can begin to identify regions of the genome and key genes that define the underlying traits of DT and reproduction in Syntrichia.
Biodiversity at different geographic scales
This interdisciplinary approach will allow us to integrate levels of biological understanding within the same field-collected genotypes: we will link patch genetics (Oliver & Fisher labs), patch physiology (Stark lab), and relationships of biodiversity (both species richness and phylogenetic diversity) to community resilience (Bowker & Mishler labs). Ultimately, this spatially and environmentally explicit framework will produce a robust, interdisciplinary picture of the functional, genetic, ecosystem, and taxonomic drivers of biodiversity in Syntrichia.